class: left, middle, inverse, title-slide # Sequence Capture of Eupolypod II Ferns:<br>Applications at Deep and Shallow Phylogenetic Levels ### Joel H. Nitta<sup>1,2</sup>, Atsushi Ebihara<sup>2</sup><br><span style="font-size: 50%;"><sup>1</sup>National Museum of Natural History, Smithsonian Institution (current)<br><sup>2</sup>National Museum of Nature and Science, Japan</span> ### 日本植物分類学会第18回大会 <br><span style="font-size: 70%;">2019.03.07</span> --- class: center, middle # なぜシダ植物は次世代<br>シーケンサーが遅れているのか? --- ## シダのゲノムは難解 *Ohioglossum reticulatum* 2*n* = 1262 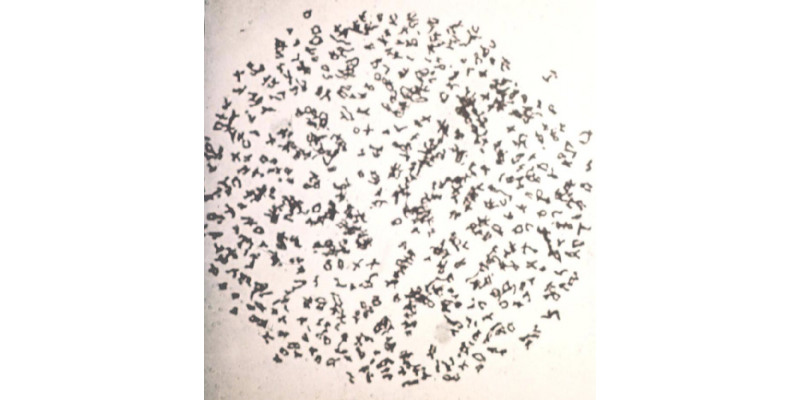 .footnote[Abraham, A. and Ninan, C.A. 1954. *L.Curr.Sci.* 23:213-214] --- ## 今までのシーケンスは葉緑体遺伝子が圧倒的に多かった  --- ## 葉緑体遺伝子による発見がいっぱい <img src="https://www.dropbox.com/s/7okeayr34ynef38/paper_pile.gif?raw=1" style="display: block; margin: auto;" /> --- class: center, middle # しかし… --- ## 葉緑体遺伝子だけでは限界がある 網状進化 <img src="index_files/figure-html/unnamed-chunk-1-1.png" width="400px" height="400px" style="display: block; margin: auto;" /> .footnote[Wagner, W. H. 1954. *Evolution* 8:103-118] --- ## 葉緑体遺伝子だけでは限界がある Ancient, rapid radiations <img src="index_files/figure-html/unnamed-chunk-2-1.png" width="600" style="display: block; margin: auto;" /> .footnote[Rothfels et al. 2012. *Sys.Bio.* 61:490-509] --- class: center, middle # やっと! --- background-position: 50% 55% background-image: url(https://ce8f5f2ec.cloudimg.io/height/400/n/https://sites.duke.edu/pryerlab/files/2018/07/nplants-Cover-JUL18_web.jpg) ## 初シダゲノム .footnote[Li, F.W. et al. 2018. *Nature Plants* 4] --- class: center, middle # シダ植物にも次世代の<br/>時代が来ました! --- class: center, middle # Methods --- ## Sequence Capture .pull-left[ シングルコピー遺伝子に絞れる 遺伝子座100以上 断片化されたDNAにも使える **ancient, rapid radiations**にも、**近縁種群**にも最適? ] .pull-right[ <img src="https://www.dropbox.com/s/hgjcyjxnhl8wg82/mybaits_seq_cap_flow.png?raw=1" height="450px" style="display: block; margin: auto;" /> ] .footnote[http://www.arborbiosci.com/] --- ## Workflow - Bait design: 37 transcriptomes → 194 genes (Yang and Smith 2014 *MBE* 31) - Library construction: NEBnext Ultra II - Sequence capture: MYBaits - Sequencing: MiSeq 2x300bp - Target recovery: HybPiper - Phylogenetic analysis: RAxML, IQ-Tree, ASTRAL <div id="htmlwidget-d854016fad4165d953f0" style="width:100%;height:302.4px;" class="grViz html-widget"></div> <script type="application/json" data-for="htmlwidget-d854016fad4165d953f0">{"x":{"diagram":"digraph {\n \n graph[layout = dot, rankdir = LR]\n labeljust=\"l\";\n \n node [shape = rectangle, style = rounded]\n a [label = \"Bait design\"]\n b [label = \"Library \n construction\"]\n c [label = \"Sequence \n capture\"]\n d [label = \"Illumina \n sequencing\"]\n e [label = \"Target \n recovery\"]\n f [label = \"Phylogenetic \n analysis\"]\n\n a -> b -> c -> d -> e -> f\n\n}","config":{"engine":"dot","options":null}},"evals":[],"jsHooks":[]}</script> --- background-position: 80% 50% background-image: url(https://www.dropbox.com/s/h2mnz4m5ojh8j2i/ppgi_japanese.png?raw=1) background-size: contain ## Materials .pull-left-narrow[ Eupolypods II<br>チャセン亜目 - ancient, rapid radiation - 11 families - ca. 2,600 spp. .footnote[Pteridophyte Phylogeny Group I. <br/> 2016. *JSE* 54:563–603.] ] --- background-position: 80% 50% background-image: url(https://www.dropbox.com/s/s2ah2clx7bfh1lk/ppgi_japanese_eupoly2.png?raw=1) background-size: contain ## Materials .pull-left-narrow[ Eupolypods II<br>チャセン亜目 - ancient, rapid radiation - 11 families - ca. 2,600 spp. .footnote[Pteridophyte Phylogeny Group I. <br/> 2016. *JSE* 54:563–603.] ] --- ## Materials .pull-left-narrow[ Eupolypods II<br>チャセン亜目 - ancient, rapid radiation - 11 families - ca. 2,600 spp. ] .pull-right-wide[ <img src="https://www.dropbox.com/s/warmnu0f6jlu4nb/ppgi_japanese_eupoly2_close.png?raw=1" width="450px" style="display: block; margin: auto 0 auto auto;" /> ] --- ## Materials .pull-left-narrow[ *Parathelypteris angustifrons* complex<br>ハシゴシダ類<br>(ヒメシダ科) - 4 taxa + hybrids - E. Asia - 2-6x ] .pull-right-wide[ <img src="https://www.dropbox.com/s/u715dd73ku8vrr3/ppgi_japanese_eupoly2_thelyp.png?raw=1" width="450px" style="display: block; margin: auto 0 auto auto;" /> ] --- background-position: 80% 50% background-image: url(https://www.dropbox.com/s/9wapy3n5yemhwqc/T_angustifrons_complex_back.png?raw=1) background-size: contain ## Materials .pull-left-narrow[ *Parathelypteris angustifrons* complex<br>ハシゴシダ類<br>(ヒメシダ科) - 4 taxa + hybrids - E. Asia - 2-6x .footnote[Ebihara, A. 2016] ] --- ## Materials サンプリング <table style="font-family:-apple-system, BlinkMacSystemFont, 'Segoe UI', Roboto, Oxygen, Ubuntu, Cantarell, 'Helvetica Neue', 'Fira Sans', 'Droid Sans', Arial, sans-serif;display:table;border-collapse:collapse;margin-left:auto;margin-right:auto;color:#000000;font-size:16px;background-color:white;width:250px;border-top-style:solid;border-top-width:2px;border-top-color:#A8A8A8; font-size: 20px; margin-left: auto; margin-right: auto;" class="table"> <tr> <th style="color:#000000;background-color:white;font-size:16px;font-weight:initial;vertical-align:middle;padding:10px;margin:10px;text-align:left;" rowspan="1" colspan="1"></th> <th style="color:#000000;background-color:white;font-size:16px;font-weight:initial;vertical-align:middle;padding:10px;margin:10px;text-align:right;font-variant-numeric:tabular-nums;" rowspan="1" colspan="1"><em>n</em></th> </tr> <tbody style="border-top-style:solid;border-top-width:2px;border-top-color:#A8A8A8;border-bottom-style:solid;border-bottom-width:2px;border-bottom-color:#A8A8A8;"> <tr style=""> <td colspan="2" style="padding:8px;color:#000000;background-color:white;font-size:16px;font-weight:initial;border-top-style:solid;border-top-width:2px;border-top-color:#A8A8A8;border-bottom-style:solid;border-bottom-width:2px;border-bottom-color:#A8A8A8;vertical-align:middle;font-size: 20px;">Eupolypods II</td> </tr> <tr> <td style="padding:10px;margin:10px;vertical-align:middle;text-align:left;background-color:white;">Herbarium specimen</td> <td style="padding:10px;margin:10px;vertical-align:middle;text-align:right;font-variant-numeric:tabular-nums;background-color:white;">4</td> </tr> <tr> <td style="background-color:#f2f2f2;padding:10px;margin:10px;vertical-align:middle;text-align:left;background-color:white;">Silica gel sample</td> <td style="background-color:#f2f2f2;padding:10px;margin:10px;vertical-align:middle;text-align:right;font-variant-numeric:tabular-nums;background-color:white;">2</td> </tr> <tr style=""> <td colspan="2" style="padding:8px;color:#000000;background-color:white;font-size:16px;font-weight:initial;border-top-style:solid;border-top-width:2px;border-top-color:#A8A8A8;border-bottom-style:solid;border-bottom-width:2px;border-bottom-color:#A8A8A8;vertical-align:middle;font-style: italic;font-size: 20px;">Parathelypteris</td> </tr> <tr> <td style="padding:10px;margin:10px;vertical-align:middle;text-align:left;background-color:white;">Herbarium specimen</td> <td style="padding:10px;margin:10px;vertical-align:middle;text-align:right;font-variant-numeric:tabular-nums;background-color:white;">6</td> </tr> <tr> <td style="background-color:#f2f2f2;padding:10px;margin:10px;vertical-align:middle;text-align:left;background-color:white;">Silica gel sample</td> <td style="background-color:#f2f2f2;padding:10px;margin:10px;vertical-align:middle;text-align:right;font-variant-numeric:tabular-nums;background-color:white;">34</td> </tr> </tbody> </table> --- class: center, middle # Results --- <img src="index_files/figure-html/efficiency-plot-1.svg" width="120%" /> --- <img src="index_files/figure-html/nuclear-recovery-plot-1.svg" width="120%" /> --- <img src="index_files/figure-html/plastid-recovery-plot-1.svg" width="120%" /> --- .small-margins[ <img src="eupoly2_plastid_tree.svg" width="120%" /> ] --- .small-margins[ <img src="eupoly2_nuc_tree.svg" width="120%" /> ] --- .small-margins[ <img src="index_files/figure-html/parathelyp-plastid-1.svg" width="120%" /> ] --- class: center, middle # Summary --- ## Sequence capture 被子植物と比べて、効率が低い - 標本から4-8% - フレッシュから10−14% - 比較:被子植物では平均27%(Johnson et al. 2018 *Syst. Bot.*) しかし、取ろうとしていた遺伝子はほとんど取れた - 標本から54% - フレッシュから97% 非ターゲットであった葉緑体遺伝子も獲得可能! --- ## Eupolypods II .pull-left-narrow[ *Desmophlebium longisorum* は、実は *Diplazium*(ヘラシダ属)だった デスモフレビウム科はmonotypicになる ] .pull-right-wide[ <img src="eupoly2_plastid_tree.svg" width="400" style="display: block; margin: auto 0 auto auto;" /> ] --- background-position: 90% 50% background-image: url(https://www.dropbox.com/s/nox1c42who7x2il/desmophleb.png?raw=1) background-size: contain ## Eupolypods II .pull-left-narrow[ *Desmophlebium longisorum* は、実は *Diplazium*(ヘラシダ属)だった デスモフレビウム科はmonotypicになる ] --- ## Eupolypods II .pull-left-narrow[ *Diplazium flavoviride* は、実は *Homalosorus*だった ] .pull-right-wide[ <img src="eupoly2_plastid_tree.svg" width="400" style="display: block; margin: auto 0 auto auto;" /> ] --- background-position: 80% 50% background-image: url(https://www.dropbox.com/s/bippgi6ju2uuotc/diplazium_flavoviride.png?raw=1) background-size: contain ## Eupolypods II .pull-left-narrow[ *Diplazium flavoviride* は、実は *Homalosorus*だった 加藤先生ズバリ ] --- background-position: 90% 50% background-image: url(https://www.dropbox.com/s/2arc0ss9fp9mres/eupoly2_summary_trees.png?raw=1) background-size: contain ## Eupolypods II .pull-left[ Rhachidosoraceae<br> (ヌリワラビ科)の<br> 位置は葉緑体と核と<br> の間で大きく変わる ] --- background-position: 50% 50% background-image: url(https://www.dropbox.com/s/upivwlpqhb8n5xy/parathelyp_summary_1.png?raw=1) background-size: contain ## *P. angustifrons* complex .pull-left[ 標本のサンプリングによって、新クレード発見 ] --- background-position: 50% 50% background-image: url(https://www.dropbox.com/s/7uczbukfqyyippf/parathelyp_summary_2.png?raw=1) background-size: contain ## *P. angustifrons* complex .pull-left[ 標本のサンプリングによって、新クレード発見 *P. grammatidoides* (フィリピン)は*P. angustifrons*の分布南限 ] --- background-position: 50% 50% background-image: url(https://www.dropbox.com/s/7uczbukfqyyippf/parathelyp_summary_2.png?raw=1) background-size: contain ## *P. angustifrons* complex .pull-left[ 標本のサンプリングによって、新クレード発見 *P. grammatidoides* (フィリピン)は*P. angustifrons*の分布南限 核遺伝子の解析はこれからです… ] --- class: center, middle # A taste of things to come # 最新研究動向 --- class: small-margins background-position: 68% 50% background-image: url(https://www.dropbox.com/s/fqtunw8spcxk76q/current_10K_fern.png?raw=1) background-size: contain .small[Kuo, L-Y, unpub.] --- class: center # 謝辞   Motomi Ito, Li-Yaung Kuo, Fay-Wei Li, Zhongyang Li, Narumi Nakato, Carl Rothfels, Xinping Qi, Haruyo Yamaguchi